using Random
using Pumas
using PumasUtilities
using AlgebraOfGraphics
using CairoMakie
using CSV
using DataFramesMeta
using Dates
PK26 - Target mediated drug disposition
1 Background
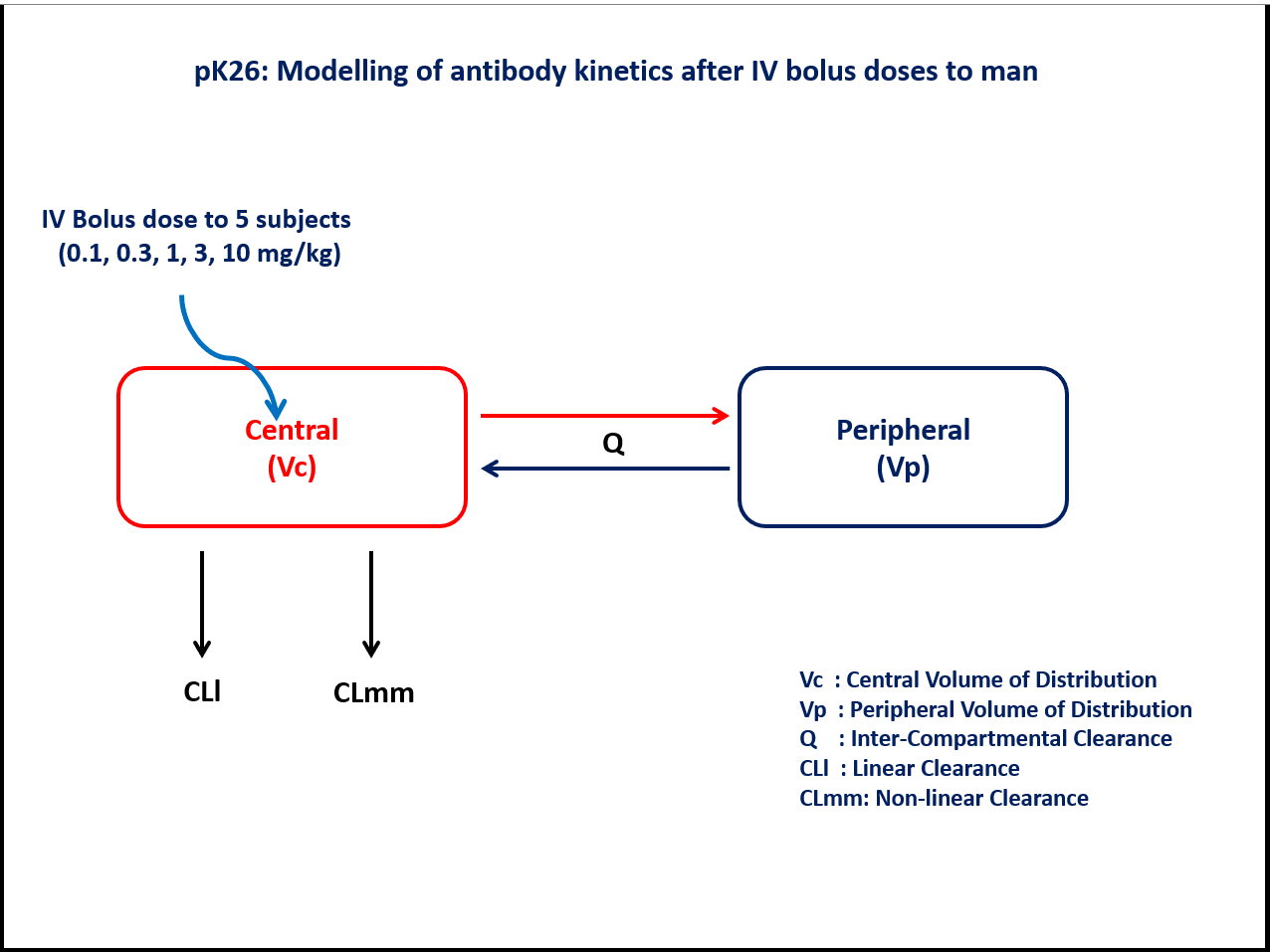
- Structural model - Two compartment model with parallel linear and non-linear elimination
- Route of administration - IV bolus (Single dose)
- Dosage Regimen - 0.1 mg/kg, 0.3 mg/kg, 1 mg/kg, 3 mg/kg, and 10 mg/kg
- Number of Subjects - 5
2 Learning Outcomes
To understand the antibody kinetics with linear and nonlinear elimination after an IV bolus dose in man.
3 Objectives
- To build a two compartment model with parallel linear and non-linear elimination to understand the antibody kinetics.
- To simulate 5 subjects after a single dose IV bolus administration
4 Libraries
Call the necessary libraries to get started.
5 Model
A two compartment model with parallel linear and non-linear elimination
pk_26 = @model begin
@metadata begin
desc = "Parallel Linear and Non-linear Elimination Model"
timeu = u"hr"
end
@param begin
"""
Maximum rate of Metabolism(mg/hr/kg)
"""
tvvmax ∈ RealDomain(lower = 0)
"""
Michaelis constant (mg/L/kg)
"""
tvkm ∈ RealDomain(lower = 0)
"""
Volume of Peripheral Compartment(L/kg)
"""
tvvp ∈ RealDomain(lower = 0)
"""
Volume of Central Compartment(L/kg)
"""
tvvc ∈ RealDomain(lower = 0)
"""
Inter-compartmental Clearance(L/hr/kg)
"""
tvq ∈ RealDomain(lower = 0)
"""
Linear Clearance(L/hr/kg)
"""
tvcll ∈ RealDomain(lower = 0)
Ω ∈ PDiagDomain(6)
"""
Proportional RUV
"""
σ²_prop ∈ RealDomain(lower = 0)
end
@random begin
η ~ MvNormal(Ω)
end
@pre begin
Vmax = tvvmax * exp(η[1])
Km = tvkm * exp(η[2])
Vp = tvvp * exp(η[3])
Vc = tvvc * exp(η[4])
Q = tvq * exp(η[5])
CLl = tvcll * exp(η[6]) # Linear clearance
# CLmm = Vmax/(Km+C) Non-linear clearance
end
@dynamics begin
Central' =
-(Vmax / (Km + (Central / Vc))) * (Central / Vc) - CLl * (Central / Vc) -
(Q / Vc) * Central + (Q / Vp) * Peripheral
Peripheral' = (Q / Vc) * Central - (Q / Vp) * Peripheral
end
@derived begin
cp = @. Central / Vc
"""
Observed Concentration (mg/L)
"""
dv ~ ProportionalNormal.(cp, sqrt(σ²_prop))
end
end┌ Warning: Variable `cp` is defined in the `@derived` block using `=` and hence `cp` is not used for model fitting but only returned when simulating: │ If `cp` is a random variable, it must be defined in the `@derived` block using `~`; │ if `cp` should be returned when simulating, it should be defined in the `@observed` block using `=`; │ if `cp` is an intermediate quantity that should not be returned when simulating, it should be defined using `:=`. └ @ Pumas ~/run/_work/PumasTutorials.jl/PumasTutorials.jl/custom_julia_depot/packages/Pumas/GZeMg/src/dsl/model_macro.jl:2351
PumasModel
Parameters: tvvmax, tvkm, tvvp, tvvc, tvq, tvcll, Ω, σ²_prop
Random effects: η
Covariates:
Dynamical system variables: Central, Peripheral
Dynamical system type: Nonlinear ODE
Derived: cp, dv
Observed: cp, dv6 Parameters
The parameters are as given below. Note that tv represents the typical value for parameters.
Vmax- Maximum rate of metabolism (mg/hr/kg)Km- Michaelis constant (mg/L/kg)Vp- Volume of Peripheral Compartment (L/kg)Vc- Volume of Central Compartment (L/kg)Q- Inter-compartmental Clearance (L/hr/kg)CLl- Linear Clearance (L/hr/kg)Ω- Between Subject Variabilityσ²_prop- Residual error
param = (
tvvmax = 0.0338,
tvkm = 0.0760,
tvvp = 0.0293,
tvvc = 0.0729,
tvq = 0.0070,
tvcll = 0.0069,
Ω = Diagonal([0.04, 0.04, 0.04, 0.04, 0.04, 0.04]),
σ²_prop = 0.04,
)7 Dosage Regimen
5 subjects received an IV bolus dose of 0.1, 0.3, 1, 3 and 10 mg/kg respectively at time=0
DR1 = DosageRegimen(0.1, time = 0)1×10 DataFrame
| Row | time | cmt | amt | evid | ii | addl | rate | duration | ss | route |
|---|---|---|---|---|---|---|---|---|---|---|
| Float64 | Int64 | Float64 | Int8 | Float64 | Int64 | Float64 | Float64 | Int8 | NCA.Route | |
| 1 | 0.0 | 1 | 0.1 | 1 | 0.0 | 0 | 0.0 | 0.0 | 0 | NullRoute |
s1 = Subject(id = "0.1 mg/kg", events = DR1, time = 0.1:0.01:1.5)Subject
ID: 0.1 mg/kg
Events: 1DR2 = DosageRegimen(0.3, time = 0)1×10 DataFrame
| Row | time | cmt | amt | evid | ii | addl | rate | duration | ss | route |
|---|---|---|---|---|---|---|---|---|---|---|
| Float64 | Int64 | Float64 | Int8 | Float64 | Int64 | Float64 | Float64 | Int8 | NCA.Route | |
| 1 | 0.0 | 1 | 0.3 | 1 | 0.0 | 0 | 0.0 | 0.0 | 0 | NullRoute |
s2 = Subject(id = "0.3 mg/kg", events = DR2, time = 0.1:0.01:7)Subject
ID: 0.3 mg/kg
Events: 1DR3 = DosageRegimen(1, time = 0)1×10 DataFrame
| Row | time | cmt | amt | evid | ii | addl | rate | duration | ss | route |
|---|---|---|---|---|---|---|---|---|---|---|
| Float64 | Int64 | Float64 | Int8 | Float64 | Int64 | Float64 | Float64 | Int8 | NCA.Route | |
| 1 | 0.0 | 1 | 1.0 | 1 | 0.0 | 0 | 0.0 | 0.0 | 0 | NullRoute |
s3 = Subject(id = "1 mg/kg", events = DR3, time = 0.1:0.1:21)Subject
ID: 1 mg/kg
Events: 1DR4 = DosageRegimen(3, time = 0)1×10 DataFrame
| Row | time | cmt | amt | evid | ii | addl | rate | duration | ss | route |
|---|---|---|---|---|---|---|---|---|---|---|
| Float64 | Int64 | Float64 | Int8 | Float64 | Int64 | Float64 | Float64 | Int8 | NCA.Route | |
| 1 | 0.0 | 1 | 3.0 | 1 | 0.0 | 0 | 0.0 | 0.0 | 0 | NullRoute |
s4 = Subject(id = "3 mg/kg", events = DR4, time = 0.1:0.1:30)Subject
ID: 3 mg/kg
Events: 1DR5 = DosageRegimen(10, time = 0)1×10 DataFrame
| Row | time | cmt | amt | evid | ii | addl | rate | duration | ss | route |
|---|---|---|---|---|---|---|---|---|---|---|
| Float64 | Int64 | Float64 | Int8 | Float64 | Int64 | Float64 | Float64 | Int8 | NCA.Route | |
| 1 | 0.0 | 1 | 10.0 | 1 | 0.0 | 0 | 0.0 | 0.0 | 0 | NullRoute |
s5 = Subject(id = "10 mg/kg", events = DR5, time = 0.1:0.1:43)Subject
ID: 10 mg/kg
Events: 1pop = [s1, s2, s3, s4, s5]Population
Subjects: 5
Observations: A more succinct way of generating the pop above across the 5 subjects is given below
doses = [0.1, 0.3, 1, 3, 10]
samp_times = [0.1:0.01:1.5, 0.1:0.01:7, 0.1:0.1:21, 0.1:0.1:30, 0.1:0.1:43]
pop = map(zip(doses, samp_times)) do (d, times)
Subject(id = string(d, " mg/kg"), events = DosageRegimen(d, time = 0), time = times)
endPopulation
Subjects: 5
Observations: 8 Simulation
To simulate plasma concentration data for 5 subjects with specific obstimes.
Random.seed!(123)The random effects are zero’ed out since we are simulating means
zfx = zero_randeffs(pk_26, pop, param)5-element Vector{@NamedTuple{η::Vector{Float64}}}:
(η = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0],)
(η = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0],)
(η = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0],)
(η = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0],)
(η = [0.0, 0.0, 0.0, 0.0, 0.0, 0.0],)sim = simobs(pk_26, pop, param, zfx)Simulated population (Vector{<:Subject})
Simulated subjects: 5
Simulated variables: cp, dv9 Visualization
@chain DataFrame(sim) begin
dropmissing(:cp)
data(_) *
mapping(
:time => "Time (days)",
:cp => "PK26 Concentrations (mg/L)",
color = :id => nonnumeric => "Doses",
) *
visual(Lines, linewidth = 4)
draw(;
axis = (
yscale = log10,
yticks = map(i -> 10.0^i, -2:2),
ytickformat = i -> (@. string(round(i; digits = 1))),
xticks = 0:10:40,
),
figure = (;
fontsize = 22,
legend = (;
linewidth = 6,
position = :bottom,
tellwidth = true,
tellheight = true,
width = Auto(),
),
),
)
end10 Population simulation
par = (
tvvmax = 0.0338,
tvkm = 0.0760,
tvvp = 0.0293,
tvvc = 0.0729,
tvq = 0.0070,
tvcll = 0.0069,
Ω = Diagonal([0.04, 0.03, 0.03, 0.02, 0.04, 0.04]),
σ²_prop = 0.04,
)
DR1 = DosageRegimen(0.1, time = 0)
DR2 = DosageRegimen(0.3, time = 0)
DR3 = DosageRegimen(1, time = 0)
DR4 = DosageRegimen(3, time = 0)
DR5 = DosageRegimen(10, time = 0)
pop1 = map(i -> Subject(id = i, events = DR1), 1:45)
pop2 = map(i -> Subject(id = i, events = DR2), 46:90)
pop3 = map(i -> Subject(id = i, events = DR3), 91:135)
pop4 = map(i -> Subject(id = i, events = DR4), 136:180)
pop5 = map(i -> Subject(id = i, events = DR5), 181:225)
pop = vcat(pop1, pop2, pop3, pop4, pop5)
Random.seed!(314)
sim_pop1 = simobs(pk_26, pop1, par, obstimes = [0.07, 0.15, 0.33, 0.43, 0.96, 1.2])
sim_pop2 = simobs(
pk_26,
pop2,
par,
obstimes = [0.23, 0.33, 0.51, 1.18, 1.75, 2.2, 3.06, 3.5, 4, 4.5, 5, 5.5, 6.89],
)
sim_pop3 = simobs(
pk_26,
pop3,
par,
obstimes = [0.2, 0.4, 1.2, 3.2, 4.5, 6, 7.1, 9, 10, 11, 12, 13, 14.3, 21],
)
sim_pop4 = simobs(
pk_26,
pop4,
par,
obstimes = [
0.4,
0.5,
0.6,
1.1,
3.2,
5,
7.1,
9,
10,
12,
14.2,
15,
16,
17,
19,
20.9,
22,
24,
25,
26,
27,
27.5,
28.1,
],
)
sim_pop5 = simobs(
pk_26,
pop5,
par,
obstimes = [
0.2,
0.5,
1.2,
2,
3.2,
5,
7.1,
9,
12,
14.1,
16,
18,
20,
21.1,
22,
24,
25,
26.5,
28.1,
30,
32,
34,
36,
38,
40,
42.1,
],
)
populationsimulation = vcat(sim_pop1, sim_pop2, sim_pop3, sim_pop4, sim_pop5)
sim_plot(populationsimulation)
df_pop = DataFrame(populationsimulation)
#CSV.write("pk_26.csv", df_pop)